Research data management (RDM)
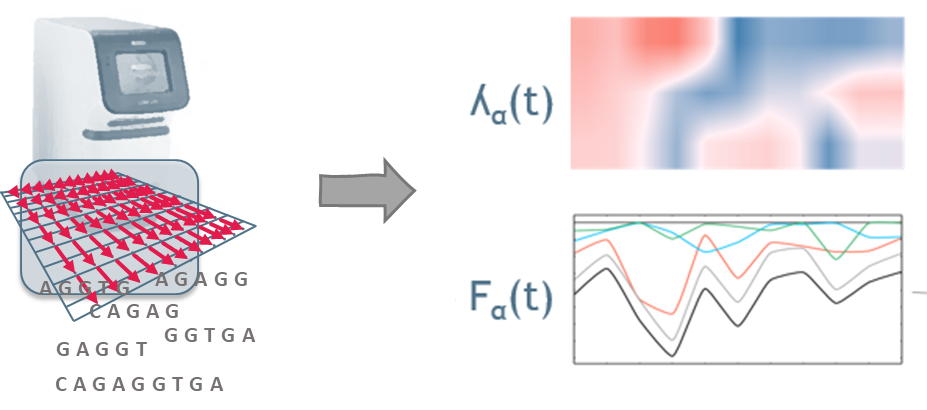
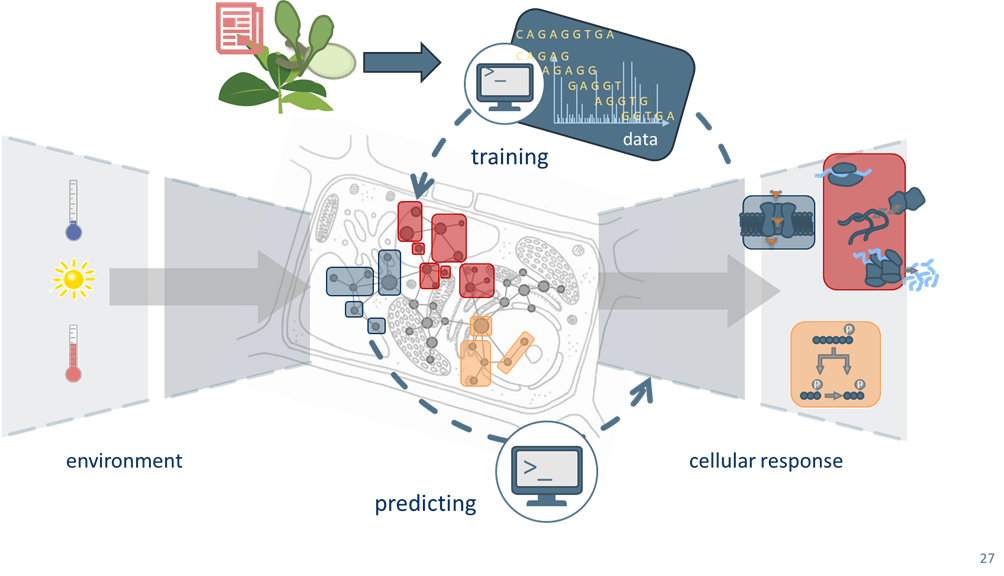
We are committed to the success of FAIR (Findable, Accessible, Interoperable, and Reusable) and open research data. Research data possess immense value, which, when combined with tomorrow's technologies such as machine and deep learning, can unlock answers to questions we cannot even conceive today. Therefore, the flexible contextualization of research data with machine-actionable metadata is essential to advance modern science.
We draw inspiration from the open-source software community's success, developing approaches that enable the biological community to collaboratively build a community-wide FAIR research data resource. Our research group participates in the National Research Data Infrastructure (NFDI) initiative with the DataPLANT project, which empowers plant researchers to engage in a thriving RDM ecosystem without barriers.